scheil
A Scheil-Gulliver simulation tool using pycalphad.
import matplotlib.pyplot as plt
from pycalphad import Database, variables as v
from scheil import simulate_scheil_solidification
# setup the simulation parameters
dbf = Database('alzn_mey.tdb')
comps = ['AL', 'ZN', 'VA']
phases = sorted(dbf.phases.keys())
liquid_phase_name = 'LIQUID'
initial_composition = {v.X('ZN'): 0.3}
start_temperature = 850
# perform the simulation
sol_res = simulate_scheil_solidification(dbf, comps, phases, initial_composition, start_temperature, step_temperature=1.0)
# plot the result
for phase_name, amounts in sol_res.cum_phase_amounts.items():
plt.plot(sol_res.temperatures, amounts, label=phase_name)
plt.plot(sol_res.temperatures, sol_res.fraction_liquid, label='LIQUID')
plt.ylabel('Phase Fraction')
plt.xlabel('Temperature (K)')
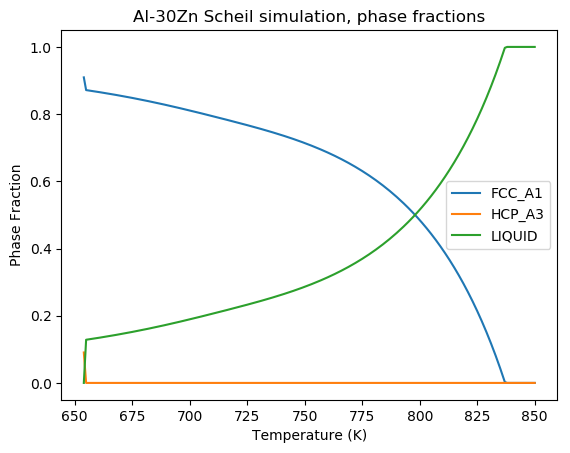
plt.title('Al-30Zn Scheil simulation, phase fractions')
plt.legend(loc='best')
plt.show()
Installation
pip (recommended)
scheil is suggested to be installed from PyPI.
pip install scheil
Anaconda
conda install -c conda-forge scheil
Development versions
To install an editable development version with pip:
git clone https://github.com/pycalphad/scheil.git
cd scheil
pip install --editable .[dev]
Upgrading scheil later requires you to run git pull in this directory.
Run the automated tests using
pytest
Theory
Uses classic Scheil-Gulliver theory (see G.H. Gulliver, J. Inst. Met. 9 (1913) 120–157 and Scheil, Zeitschrift Für Met. 34 (1942) 70–72.) with assumptions of
- Perfect mixing in the liquid
- Local equilibrium between solid and liquid
- No diffusion in the solid
Getting Help
For help on installing and using scheil, please join the pycalphad/pycalphad Gitter room.
Bugs and software issues should be reported on GitHub.
License
scheil is MIT licensed. See LICENSE.
Citing
If you use the scheil package in your work, please cite the relevant version.
The following DOI, doi:10.5281/zenodo.3630656, will link to the latest released version of the code on Zenodo where you can cite the specific version that you haved used. For example, version 0.1.2 can be cited as:
Bocklund, Brandon, Bobbio, Lourdes D., Otis, Richard A., Beese, Allison M., & Liu, Zi-Kui. (2020, January 29). pycalphad-scheil: 0.1.2 (Version 0.1.2). Zenodo. http://doi.org/10.5281/zenodo.3630657
@software{bocklund_brandon_2020_3630657,
author = {Bocklund, Brandon and
Bobbio, Lourdes D. and
Otis, Richard A. and
Beese, Allison M. and
Liu, Zi-Kui},
title = {pycalphad-scheil: 0.1.2},
month = jan,
year = 2020,
publisher = {Zenodo},
version = {0.1.2},
doi = {10.5281/zenodo.3630657},
url = {https://doi.org/10.5281/zenodo.3630657}
}